[ Spring 2008 ]
A math whiz takes on brain cancer, MS, and Alzheimer’s disease.
Rebecca Betensky’s dad worked as a statistician for a global oil company on “credit rating stuff,” she says, so for a long time, she “stayed as far away from statistics as I could.” She had settled on a career in theoretical mathematics when, as a junior at Harvard College in 1985, she had a change of heart.
Studying human physiology and epidemiology with Joseph Brain, a professor at the Harvard School of Public Health (HSPH), showed her that, “Wow—you can apply math to all these interesting medical questions.” A chance to work for a year with biostatisticians on patient studies at Boston’s Dana-Farber Cancer Institute after graduation was the clincher.
Ever since then, Betensky has been inventing new methods of capturing, interpreting, and using patient data, helping doctors tailor patients’ therapies to the special characteristics of their condition. With specialists at Boston’s teaching hospitals, she now focuses on neurological diseases—brain cancers, multiple sclerosis, Alzheimer’s disease.
No matter the illness, “The questions are often the same,” Betensky says. “How can we diagnose patients in a way that reflects the course of their disease? Can we do it more accurately than by looking at their cells under a microscope, or at their current symptoms and test results?”
In this new century, it is possible to stratify patients according to their unique genetic traits. Advanced technologies can mine so much information from a patient’s DNA that making sense of it is an Everest climb. “The sheer volume of data is the challenge—it’s a very big problem, actually,” says Betensky, who came to HSPH in 1994 and was named a professor of biostatistics last year.
The Holy Grail is to sort patients into subclasses of prognoses, then come up with customized treatments that ideally would be targeted to their own DNA. Patients with poor prognoses might choose an aggressive experimental therapy—or perhaps focus on end-of-life issues. A brighter outlook, on the other hand, would bring relief, and many more precious years.
Same diagnosis, different outlook
Take the cancer known as oligodendroglioma. As brain tumors go, “This one is a ‘good’ kind to have, because you might live a long time,” Betensky says. Nearly 11,000 people in the United States annually receive this diagnosis, yet their fates are anything but uniform. Half live longer than 10 years. The rest die sooner, in as little as two years.
No More Math AnxietyHSPH’s 2007 Quantitative Sciences program, led by (far right) instructor Andy Houseman, director Rebecca Betensky, and (front row, second from left) coordinator Catherine Haskell, included college students from across the U.S., all potential innovators in biostatistics and epidemiology. “A role model for young women statisticians” is what the Harvard Medical School’s Dianne Finkelstein called Rebecca Betensky in a 2005 letter nominating her for the American Public Health Association’s top prize for biostatisticians under age 40, the Mortimer Speigelman Award. Betensky won the prize, but Finkelstein was only half right. “I guess I’m called a role model for women because I’ve managed to become a Harvard professor while raising three children,” Betensky concedes with a chuckle. For that, she gives her husband partial credit. But when it comes to mentoring students, Betensky takes all talented comers. As head of the Initiative for Maximizing Student Diversity since 2003, she’s advised more than two dozen doctoral candidates, male and female. Several are now professors in their own right. The program, launched 15 years ago by HSPH Department of Biostatistics Chair Louise Ryan, continues the department’s well-established tradition of attracting minorities and women. Undergraduate students, too, train under Betensky’s wing. Since 2003, she has co-directed a month-long summer program at HSPH that draws minority and first-generation college students to her field from around the country. “We measure our success by those who go on to graduate school—or even better, come to HSPH,” Betensky notes with satisfaction. “We’re working very hard to keep the pipeline filled.” |
Which patients are which? Looking for answers, David Louis, the chief of pathology at Boston’s Massachusetts General Hospital, turned to Betensky. For her PhD thesis at Stanford, she had come up with a methodology to monitor patients’ progress in clinical trials that compared three or more treatment groups, allowing for stopping a trial early if one treatment appeared to be highly effective. Later, her calculations helped show researchers when to stop a trial early in cases where an experimental therapy proved futile. And, in a large study of patients with two kinds of cancer—breast and ovarian, both of which can run in families—she had devised a new way to estimate the risks of these diseases, after first taking into account patients’ own and family histories and the criteria by which families were selected for study.
At MGH, Louis had combed the chromosomes of 93 oligodendroglioma patients for clues to their prognosis. Like all humans, each patient had 23 pairs made up of matching DNA strands, with each chromosome having two “arms.” Louis had discovered that as many as 60 percent of tumors were missing DNA from entire arms of chromosomes 1 and 19. His lab tested for losses at seven locations, or “markers,” along each strand. If DNA was missing from informative, or “heterozygous,” sites—a condition known to oncologists as “loss of heterozygosity,” or LOH—it meant that key but as yet unidentified genes were lost. Mysteriously, patients with LOH did much better on chemotherapy than the rest.
How exactly were LOH status and prognosis related? Louis and his collaborators decided to test patients at additional markers—19 in all. Then he gave Betensky his data. “Too many variables and not enough patients” was her quandary, she remembers.
In a flash of insight, she and two colleagues dusted off a technique from 1950s education and psychology research. This strategy, known as latent-class modeling, grouped patients into useful categories based on just a few strategically chosen variables. For example, the scale of clinical depression ranks subjects from one to five. Disability among the elderly is assessed according to one’s ability to perform five tasks of varying degrees of difficultly, including walking and climbing stairs.
‘Elegant’ solution
Instead of assigning equal weight to all variables, the trio developed a statistical algorithm that stressed the significance of certain variables over others. Their innovation was, first, to combine scientists’ knowledge that losing the tip of chromosome 1 was associated with a good prognosis with their uncertainty as to what losing other regions might mean; and second, to let the data sort itself into subgroups.
“In other words, we set up our model to give more weight to certain regions of the chromosomes, much as you might handicap horses in a race by giving certain ones a head start—the tallest and most common breeds, say,” Betensky explains. “We knew the tip of chromosome 1 would be an important distinguishing factor among patients, so we gave markers there the ‘head start.’ However, if a subgroup of patients had a consistent pattern of loss of heterozygosity at chromosome 1’s less-explored middle region, our algorithm would also identify that region as an important classification factor.”
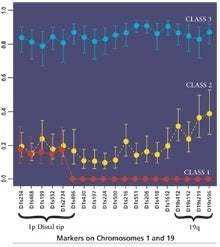 By the time they finished, Betensky and former postdoctoral fellow Andy Houseman, now an assistant professor at University of Massachusetts-Lowell, and Brent Coull, an associate professor at HSPH, had sorted 93 patients into three distinct risk groups based on LOH status. Classes 1 and 2 would fare poorly, while Class 3 had a far brighter prognosis (see graph).
By the time they finished, Betensky and former postdoctoral fellow Andy Houseman, now an assistant professor at University of Massachusetts-Lowell, and Brent Coull, an associate professor at HSPH, had sorted 93 patients into three distinct risk groups based on LOH status. Classes 1 and 2 would fare poorly, while Class 3 had a far brighter prognosis (see graph).
A colleague of Betensky’s, Karen Bandeen-Roche, a professor of biostatistics at the Johns Hopkins Bloomberg School of Public Health, admires this inventive strategy, which awaits validation using data from larger numbers of patients. When researchers don’t have enough information to partition a population precisely, “The best they can do is infer subcategories from surrogate measures, such as disease symptoms,” Bandeen-Roche says. One alternative is to simply set your sorting criteria, “but then you might get an answer that’s inaccurate due to oversimplification,” she cautions. Betensky’s team’s method “elegantly compromises between the two extremes.”
In David Louis’ view, Betensky is a breed apart. “Some biostatisticians approach data analysis as if it were a task,” he says. “Rebecca studies the data to find some new way of mining it for information, in ways no one’s ever done before.”
Predicting MS
Betensky has also applied her statistical sleight of hand to multiple sclerosis, a disease whose symptoms and course are highly variable. For years, neurologists have monitored the disease using the Extended Disability Status Scale,
or EDSS. Patients are annually assigned a value of 1 to 9 to reflect their coordination, degree of movement, vision, speech, loss of sensation, and other signs of disease progression.
A few years ago, through her role as biostatistics director of Harvard’s NeuroDiscovery Center, a consortium of hospitals affiliated with Harvard’s medical school, Betensky and post-doctoral student Micha Mandel were approached by Howard Weiner, the director of the Partners Multiple Sclerosis Center at Brigham and Women’s Hospital, who asked: Could EDSS data be used to predict a patient’s future course?
“What the doctors said was, ‘If someone walks in the door and they’re at 3 today, and this is their history, what’s the chance they’ll be at a 6, or in a wheelchair, in three years?’” The challenge was tricky. One problem, Betensky says, is that the EDSS isn’t evenly calibrated: “Progressing from 1 to 2, say, is less onerous than the shift from 3 to 4.” Also, since patients repeatedly go in and out of remission, their values fluctuate over time—3, 3, 4, 6, 5, for example. Moreover, MS symptoms don’t progress in lock-step; one might remain mild while another grows severe.
In 2007, the HSPH statisticians published their analysis, showing the probability of disease progression given patients’ EDSS history. Starting with a limited pool of patient data from the MS Center at Brigham and Women’s Hospital, they added in MRI and other test results.
As more data come in, they will perfect their methodology and invite others to validate it. Their hope is to one day create a web-based software program useful for planning patients’ therapy, as well as their futures.
Identifying Alzheimer’s early
Rather than make do with what data she’s given, Betensky prefers to get in on the ground floor of designing clinical trials. Opportunities abound. Take Alzheimer’s disease, for which there is neither preventive nor cure.
At MGH, where Betensky is a member of the Department of Medicine, she and psychiatrist Deborah Blacker have had some success identifying early-stage patients who are likely to progress quickly and are therefore ideal candidates for trials. “The thought is, any new treatment will most likely work on patients very early in the course of their disease. If you expect someone to progress quickly, you’d also expect them to be an ideal clinical trial participant, since either a new treatment will slow their expected progression, or it won’t.”
The researchers are looking for useful early warning flags by following patients who initially sought an evaluation from the hospital’s memory disorders clinic. “We’re asking, ‘Which of these can we use as clinical endpoints in a trial involving very-early-stage patients, to know whether or not a treatment is helping?’ ” It may turn out that patients for whom a standardized test shows a high rate of cognitive decline within just three years of troubling memory lapses are destined to get Alzheimer’s, and so would be good candidates for trials of new therapies.
Too much information?
Asked what other biostatistical brain teasers she’ll tackle next, Betensky returns to the topic of brain cancer. Building on what she’s learned from genetic analyses of oligodendrogliomas, she’ll be working with David Louis on the uniformly lethal gliomas as well as the meningiomas, most of which are benign (non-spreading). Relatively few in this latter group are decidedly malignant; the rest fall into a murky, “atypical” category. Which tumors within this gray area will have bad prognoses?
“We’re looking at atypicals to see if the genetics can tell us something useful about these patients’ futures,” Betensky says. In cancer, the most promising new treatments are targeted to specific genes, she points out. Low-risk patients could rejoice. Those judged high-risk might opt for surgery, or perhaps an experimental treatment.
Thanks to a technological advance known as array comparative genomic hybridization, Louis and Betensky’s meningioma studies are examining hundreds of thousands of markers. The result is a staggering number of genetic variables for every person studied.
It’s enough to boggle the mind, even a statistician’s. “How do we scale methods we’ve managed to develop, first for 19 and then for 1,000 markers, up to 100,000?” Betensky asks with amazement.
“If you look at the 2007 and 2008 literature, there’s still no consensus on how to handle all that data—even right here, among several groups working on this problem at Harvard.” At workshops and conferences, Betensky enjoys pulling experts together, figuring two or more heads will be better than one.
Karin Kiewra is editor of the Review and Associate Director of Development Communications
Portrait: Kent Dayton/HSPH. Background: ©Francois Paquet-Durand/Photo Researchers, Inc.

