Presented by: Yered Pita-Juarez
Authored by: Yered Pita-Juarez, Liuliu Pan, Jonathan Hecht, Tyler Heather, Linus Tsai, Yury Popov, Jason Reeves, Pourya Naderi, Stefan Riedel, Eric Miller, Winston Hide, Sarah Warren, Gyongyi Szabo, Joseph Beechem, Z. Gordon Jiang, Ioannis S. Vlachos
If you have any questions regarding the poster, feel free to reach out to Yered Pita-Juarez on Slack or email.
Abstract
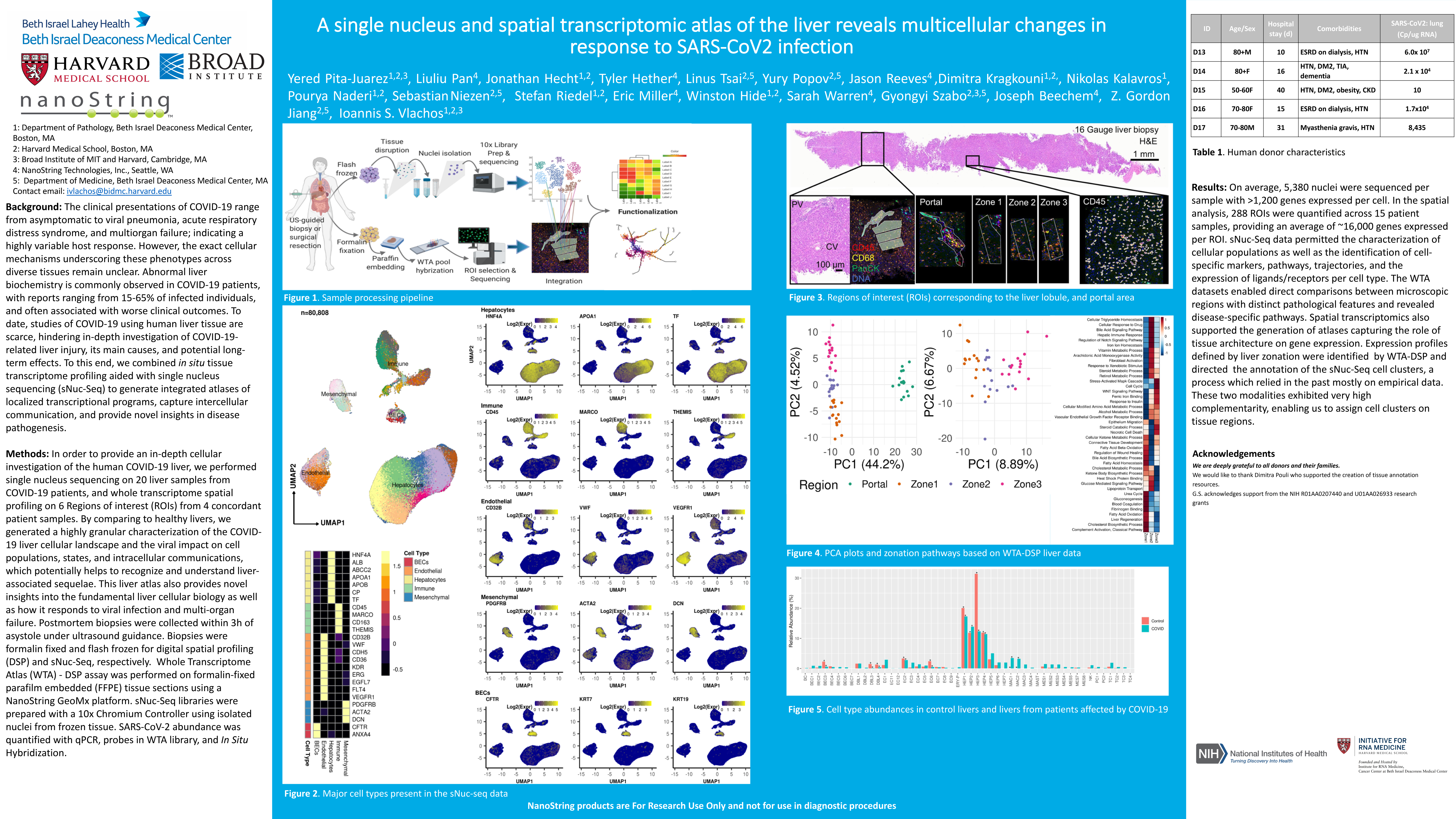
COVID-19 exhibits a wide phenotypic spectrum with potential multi-organ involvement during its acute phase. Abnormal liver biochemistry is commonly observed in COVID-19 patients, with reports ranging from 15-65% of infected individuals, and often associated with worse clinical outcomes. To date, studies of COVID-19 using human liver tissue are scarce, hindering in-depth investigation of COVID-19-related liver injury, its main causes, and potential long-term effects.
In order to provide an in-depth cellular investigation of the human COVID-19 liver, we performed single nucleus sequencing on 20 liver samples from COVID-19 patients, and whole transcriptome spatial profiling on 6 Regions of interest (ROIs) from 4 concordant patient samples. By comparing to healthy livers, we generated a highly granular characterization of the COVID-19 liver cellular landscape and the viral impact on cell populations, states, and intracellular communications, which potentially helps to recognize and understand liver-associated sequelae. This liver atlas also provides novel insights into the fundamental liver cellular biology as well as how it responds to viral infection and multi-organ failure.